Simulating properties of elastin-like polypeptides
A collaboration with Gijsje Koenderink
A biomaterial is a material specifically engineered to interact with biological systems, particularly cells and tissues. Biomaterials hold great potential for medical and nutritional purposes. However, currently biomaterials often rely on animal derived proteins, which limits their tunability and raises environmental and societal concerns. We aim to sustainably produce both natural and engineered structural proteins, notably (natural, full length) collagen and elastin, the two main building blocks of the mammalian extracellular matrix (ECM). With these proteins we aim to construct a biomaterial with cell-instructive cues and test them as cell scaffolds for soft tissue repair and cell-based meat production.
Collagen is organized into large fibres and networks and contributes to tissue strength, flexibility and support. Elastin, on the other hand, provides tissue with elasticity and resilience. In order to mimic the unique properties of elastin, scientists developed de novo elastin-like polypeptides (ELPs). One very unique property of ELPs is the so-called “lower critical solvation temperature” (LCST), this the temperature (Tt) at which they transition from being soluble in water to insoluble, driven by hydrophobic interactions that cause them to aggregate and phase separate. A Tt at physiological conditions (37˚C) is crucial for ELPs used in biomedical application. However, the value of Tt depends on numerous variables, such as type and placement of amino acid, linkers and chain length, and is therefore unknown for de novo ELPs.
In this computational project, we aim to simulate different novel ELP designs with molecular dynamics simulations (MD). In these ELP designs, we can play with the type and placement of amino acids, introducing certain linker blocks, the molecular weight, and buffer conditions.
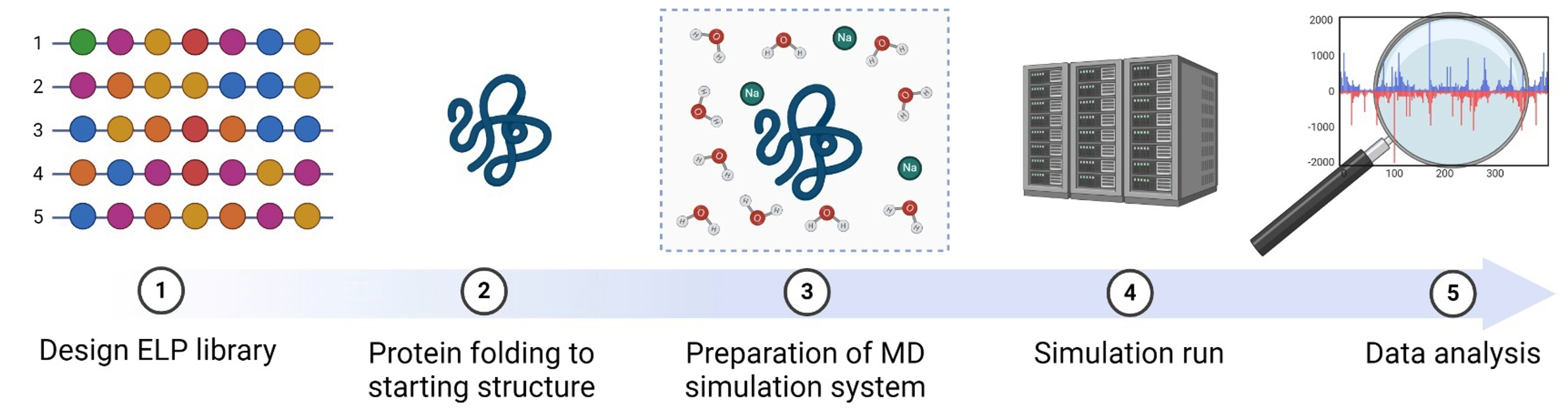
The project consists of two main parts, constructing the pipeline for the MD simulations (step 2 and 3 in the figure), and, after running the simulations (step 4), analysing the data (step 5). The ELP with the most promising predicted properties will be used for further experimental research in the future.
During this project, you will gain experience with MD simulations and develop a better understanding of protein structure and function and data analysis skills.
The project will be co-supervised by Christine Visser.